
| Gene: Otud5 | ID: uc009smt.1_intron_3_0_chrX_7444950_f | SPECIES: mm9 |
 |
 |
|
(3) OTHER.mut |
(5) PIWI.ip |
(2) PIWI.mut |
(20) TESTES |
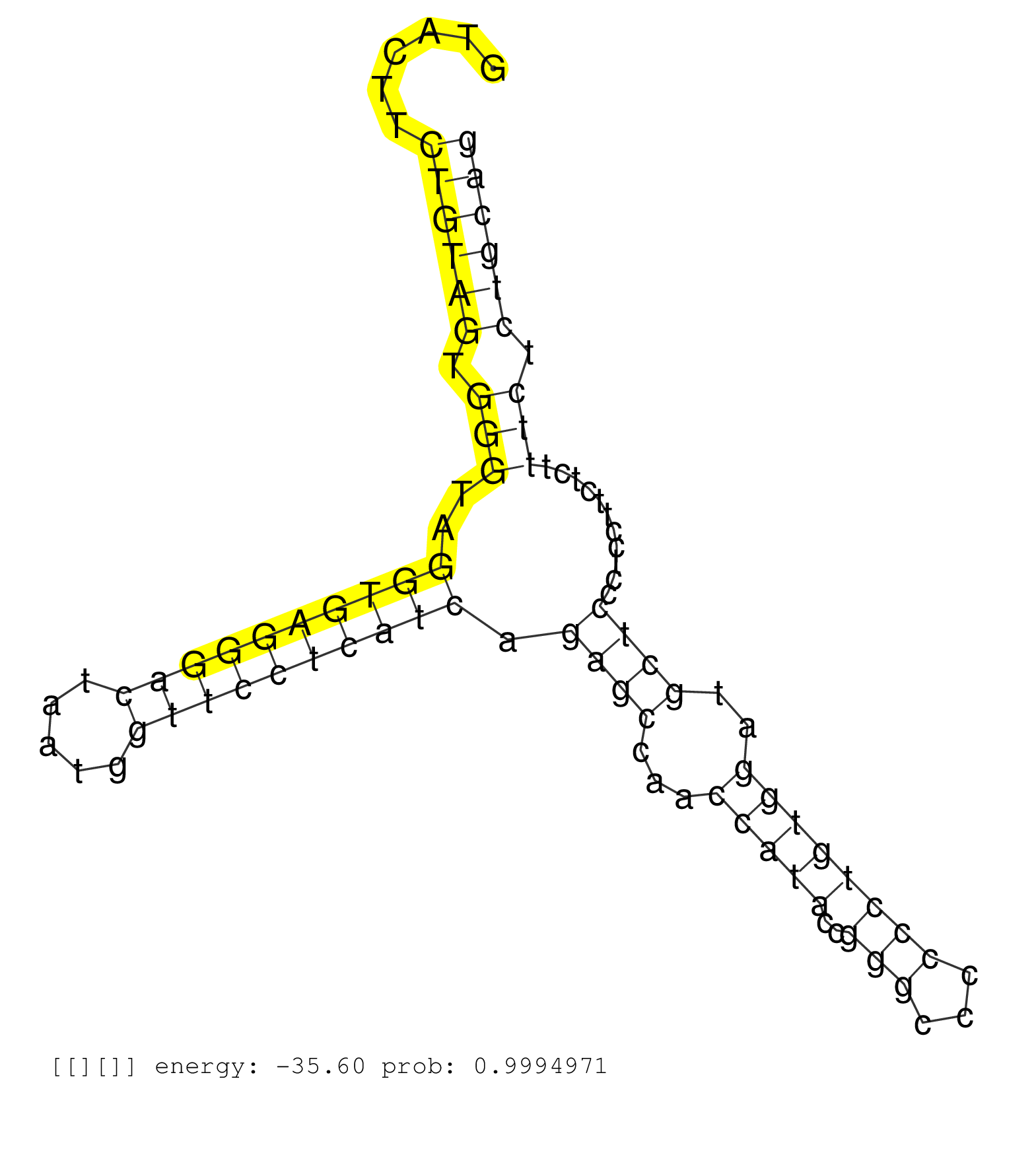
| TATGGCAGAGATGTACAACCGTCCTGTGGAGGTGTATCAATATAGCACAGGTACTTCTGTAGTGGGTAGGTGAGGGACTAATGGTTCCTCATCAGAGCCAACCATACCGGGCCCCCCTGTGGATGCTCCCTCCTTCTCTTTCTCTGCAGAACCTATCAACACATTCCATGGGATCCATCAAAATGAAGATGAACCCATC ........................................................((((((.(((..((((((((((.....)))))))))).((((...(((((..(((...))))))))..))))...........))).)))))).................................................. ..................................................51................................................................................................149................................................ |
Size | Perfect hit | Total Norm | Perfect Norm | SRR034118(GSM466728) Mili IP_Tdrd9+/- replicate1. (mili testes) | SRR034119(GSM466729) Mili IP_Tdrd9+/- replicate2. (mili testes) | GSM509280(GSM509280) small RNA cloning by length. (testes) | SRR028732(GSM400969) Mili-Tdrd1 KO associated. (mili testes) | SRR069809(GSM610965) small RNA sequencing; sample 1. (testes) | SRR034121(GSM466731) Mili IP_Tdrd9-/- replicate2. (mili testes) | SRR248523(GSM733811) cell type: Thy1+ spermatogonial stem cellstra. (testes) | GSM509277(GSM509277) small RNA cloning by length. (piwi testes) | mjTestesKO7() Testes Data. (Zcchc11 testes) | SRR248524(GSM733812) cell type: Thy1+ spermatogonial stem cellstra. (testes) | SRR034120(GSM466730) Mili IP_Tdrd9-/- replicate1. (mili testes) | SRR029042(GSM433294) 18.5dpc_hetero_tdrd1-KO. (tdrd1 testes) | mjTestesKO8() Testes Data. (Zcchc11 testes) | SRR248527(GSM733815) cell type: spermatogonial stem cell enriched . (testes) | SRR248526(GSM733814) cell type: Thy1- spermatogonial stem cellstra. (testes) | SRR014231(GSM319955) 16.5 dpc total. (testes) | SRR248525(GSM733813) cell type: Thy1- spermatogonial stem cellstra. (testes) | GSM509276(GSM509276) small RNA cloning by length. (testes) | GSM509279(GSM509279) MVH-/- E16.5 small RNA. (testes) | GSM475280(GSM475280) Mili-IP. (mili testes) |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| ..................................................GTACTTCTGTAGTGGGTAGGTGAGGG........................................................................................................................... | 26 | 1 | 9.00 | 9.00 | 1.00 | 2.00 | - | 1.00 | - | 2.00 | 2.00 | - | - | 1.00 | - | - | - | - | - | - | - | - | - | - |
| ..................................................GTACTTCTGTAGTGGGTAGGTGAGG............................................................................................................................ | 25 | 1 | 7.00 | 7.00 | - | 1.00 | - | - | - | - | 2.00 | 1.00 | 2.00 | 1.00 | - | - | - | - | - | - | - | - | - | - |
| ..................................................GTACTTCTGTAGTGGGTAGGTGAGGGt.......................................................................................................................... | 27 | t | 7.00 | 9.00 | 1.00 | 1.00 | - | 1.00 | - | 2.00 | - | - | - | 1.00 | 1.00 | - | - | - | - | - | - | - | - | - |
| ................AACCGTCCTGTGGAGGTGTATCAATAT............................................................................................................................................................ | 27 | 1 | 6.00 | 6.00 | 1.00 | 1.00 | 1.00 | 1.00 | - | 1.00 | - | - | - | - | 1.00 | - | - | - | - | - | - | - | - | - |
| ..............ACAACCGTCCTGTGGAGGTGTATCAAT.............................................................................................................................................................. | 27 | 1 | 3.00 | 3.00 | 3.00 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - |
| ................AACCGTCCTGTGGAGGTGTATCAATA............................................................................................................................................................. | 26 | 1 | 3.00 | 3.00 | 1.00 | - | - | 1.00 | - | - | - | - | - | - | 1.00 | - | - | - | - | - | - | - | - | - |
| .................ACCGTCCTGTGGAGGTGTATCAATATA........................................................................................................................................................... | 27 | 1 | 2.00 | 2.00 | - | - | 1.00 | - | - | - | - | 1.00 | - | - | - | - | - | - | - | - | - | - | - | - |
| ..................................................GTACTTCTGTAGTGGGTAGGTGAGGt........................................................................................................................... | 26 | t | 2.00 | 7.00 | 1.00 | 1.00 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - |
| ...................................................TACTTCTGTAGTGGGTAGGTGAGGGA.......................................................................................................................... | 26 | 1 | 2.00 | 2.00 | - | - | - | - | 1.00 | - | - | 1.00 | - | - | - | - | - | - | - | - | - | - | - | - |
| ........AGATGTACAACCGTCCTGTGGAGGTGT.................................................................................................................................................................... | 27 | 1 | 2.00 | 2.00 | - | 1.00 | - | - | - | - | - | - | - | - | - | - | - | - | - | 1.00 | - | - | - | - |
| ..................................................GTACTTCTGTAGTGGGTAGGTGAGGGA.......................................................................................................................... | 27 | 1 | 2.00 | 2.00 | - | - | - | - | - | - | - | 1.00 | - | - | - | - | - | - | - | - | - | - | 1.00 | - |
| ..............ACAACCGTCCTGTGGAGGTGTATCAATAT............................................................................................................................................................ | 29 | 1 | 2.00 | 2.00 | - | - | 1.00 | 1.00 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - |
| ......................................................TTCTGTAGTGGGTAGGTGAGGGACT........................................................................................................................ | 25 | 1 | 1.00 | 1.00 | - | 1.00 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - |
| .................ACCGTCCTGTGGAGGTGTATCAATATAG.......................................................................................................................................................... | 28 | 1 | 1.00 | 1.00 | - | - | 1.00 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - |
| .........................................................TGTAGTGGGTAGGTGAGGG........................................................................................................................... | 19 | 1 | 1.00 | 1.00 | - | - | - | - | - | - | - | - | - | - | - | 1.00 | - | - | - | - | - | - | - | - |
| ...................CGTCCTGTGGAGGTGTATCAATt............................................................................................................................................................. | 23 | t | 1.00 | 0.00 | - | - | - | - | 1.00 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - |
| ................AACCGTCCTGTGGAGGTGTATCAATATAG.......................................................................................................................................................... | 29 | 1 | 1.00 | 1.00 | - | - | 1.00 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - |
| ..................CCGTCCTGTGGAGGTGTATCAATATAGCA........................................................................................................................................................ | 29 | 1 | 1.00 | 1.00 | - | - | - | - | - | - | - | 1.00 | - | - | - | - | - | - | - | - | - | - | - | - |
| ..............................................................TGGGTAGGTGAGGGACTAATGGTT................................................................................................................. | 24 | 1 | 1.00 | 1.00 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | 1.00 | - | - | - |
| ..................CCGTCCTGTGGAGGTGTATCAA............................................................................................................................................................... | 22 | 1 | 1.00 | 1.00 | - | - | - | - | 1.00 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - |
| .............TACAACCGTCCTGTGGAGGTGT.................................................................................................................................................................... | 22 | 1 | 1.00 | 1.00 | - | - | - | - | 1.00 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - |
| .......GAGATGTACAACCGTCCTGTGGAGGTGT.................................................................................................................................................................... | 28 | 1 | 1.00 | 1.00 | 1.00 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - |
| .................................GTATCAATATAGCACAG..................................................................................................................................................... | 17 | 1 | 1.00 | 1.00 | - | - | - | - | - | - | - | - | - | - | - | - | - | 1.00 | - | - | - | - | - | - |
| ..................................................GTACTTCTGTAGTGGGTAGGTG............................................................................................................................... | 22 | 1 | 1.00 | 1.00 | - | - | - | - | - | - | - | - | 1.00 | - | - | - | - | - | - | - | - | - | - | - |
| ...................................................TACTTCTGTAGTGGGTAGGTGAGGG........................................................................................................................... | 25 | 1 | 1.00 | 1.00 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | 1.00 | - | - | - | - | - |
| ...........TGTACAACCGTCCTGTGGAGGTGTATC................................................................................................................................................................. | 27 | 1 | 1.00 | 1.00 | - | - | - | 1.00 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - |
| .................ACCGTCCTGTGGAGGTGTATCA................................................................................................................................................................ | 22 | 1 | 1.00 | 1.00 | - | - | 1.00 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - |
| .........GATGTACAACCGTCCTGTGGA......................................................................................................................................................................... | 21 | 1 | 1.00 | 1.00 | - | - | - | - | - | - | - | - | 1.00 | - | - | - | - | - | - | - | - | - | - | - |
| ....................................................................................................................................................................TCCATGGGATCCATCAAAATGAAGATGA....... | 28 | 1 | 1.00 | 1.00 | - | - | - | - | - | - | - | - | - | 1.00 | - | - | - | - | - | - | - | - | - | - |
| ..............ACAACCGTCCTGTGGAGGTGTATCAATA............................................................................................................................................................. | 28 | 1 | 1.00 | 1.00 | 1.00 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - |
| .................ACCGTCCTGTGGAGGTGTAT.................................................................................................................................................................. | 20 | 1 | 1.00 | 1.00 | - | - | 1.00 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - |
| ................AACCGTCCTGTGGAGGTGTATCAATATt........................................................................................................................................................... | 28 | t | 1.00 | 6.00 | - | - | - | 1.00 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - |
| .................ACCGTCCTGTGGAGGTGTATCAATAT............................................................................................................................................................ | 26 | 1 | 1.00 | 1.00 | - | 1.00 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - |
| ..................................................GTACTTCTGTAGTGGGTAGGTGA.............................................................................................................................. | 23 | 1 | 1.00 | 1.00 | - | - | - | - | - | - | - | - | - | - | - | - | 1.00 | - | - | - | - | - | - | - |
| .....................TCCTGTGGAGGTGTATCAATATAGCACAG..................................................................................................................................................... | 29 | 1 | 1.00 | 1.00 | - | - | 1.00 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - |
| ..................................................GTACTTCTGTAGTGGGTAGGTGAGGGtt......................................................................................................................... | 28 | tt | 1.00 | 9.00 | - | - | - | - | - | - | - | - | - | - | 1.00 | - | - | - | - | - | - | - | - | - |
| ...........TGTACAACCGTCCTGTGGAGGTGTATCt................................................................................................................................................................ | 28 | t | 1.00 | 1.00 | 1.00 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - |
| .............TACAACCGTCCTGTGGAGGggt.................................................................................................................................................................... | 22 | ggt | 1.00 | 0.00 | - | - | - | - | 1.00 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - |
| ..............ACAACCGTCCTGTGGAGGTG..................................................................................................................................................................... | 20 | 1 | 1.00 | 1.00 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | 1.00 | - | - |
| ...............CAACCGTCCTGTGGAGGTGTA................................................................................................................................................................... | 21 | 1 | 1.00 | 1.00 | - | - | - | - | 1.00 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - |
| .............TACAACCGTCCTGTGGAGGTGTATCAAT.............................................................................................................................................................. | 28 | 1 | 1.00 | 1.00 | - | 1.00 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - |
| ..............ACAACCGTCCTGTGGAGGTGTATCAA............................................................................................................................................................... | 26 | 1 | 1.00 | 1.00 | 1.00 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - |
| ...................................................................................................................................................................................AAAATGAAGATGAACCCA.. | 18 | 2 | 0.50 | 0.50 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | 0.50 |
| TATGGCAGAGATGTACAACCGTCCTGTGGAGGTGTATCAATATAGCACAGGTACTTCTGTAGTGGGTAGGTGAGGGACTAATGGTTCCTCATCAGAGCCAACCATACCGGGCCCCCCTGTGGATGCTCCCTCCTTCTCTTTCTCTGCAGAACCTATCAACACATTCCATGGGATCCATCAAAATGAAGATGAACCCATC ........................................................((((((.(((..((((((((((.....)))))))))).((((...(((((..(((...))))))))..))))...........))).)))))).................................................. ..................................................51................................................................................................149................................................ |
Size | Perfect hit | Total Norm | Perfect Norm | SRR034118(GSM466728) Mili IP_Tdrd9+/- replicate1. (mili testes) | SRR034119(GSM466729) Mili IP_Tdrd9+/- replicate2. (mili testes) | GSM509280(GSM509280) small RNA cloning by length. (testes) | SRR028732(GSM400969) Mili-Tdrd1 KO associated. (mili testes) | SRR069809(GSM610965) small RNA sequencing; sample 1. (testes) | SRR034121(GSM466731) Mili IP_Tdrd9-/- replicate2. (mili testes) | SRR248523(GSM733811) cell type: Thy1+ spermatogonial stem cellstra. (testes) | GSM509277(GSM509277) small RNA cloning by length. (piwi testes) | mjTestesKO7() Testes Data. (Zcchc11 testes) | SRR248524(GSM733812) cell type: Thy1+ spermatogonial stem cellstra. (testes) | SRR034120(GSM466730) Mili IP_Tdrd9-/- replicate1. (mili testes) | SRR029042(GSM433294) 18.5dpc_hetero_tdrd1-KO. (tdrd1 testes) | mjTestesKO8() Testes Data. (Zcchc11 testes) | SRR248527(GSM733815) cell type: spermatogonial stem cell enriched . (testes) | SRR248526(GSM733814) cell type: Thy1- spermatogonial stem cellstra. (testes) | SRR014231(GSM319955) 16.5 dpc total. (testes) | SRR248525(GSM733813) cell type: Thy1- spermatogonial stem cellstra. (testes) | GSM509276(GSM509276) small RNA cloning by length. (testes) | GSM509279(GSM509279) MVH-/- E16.5 small RNA. (testes) | GSM475280(GSM475280) Mili-IP. (mili testes) |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| .................................................................................................................................CTCCTTCTCTTTCTCTGCAGAACCTA............................................ | 26 | 1 | 1.00 | 1.00 | - | - | - | - | - | - | 1.00 | - | - | - | - | - | - | - | - | - | - | - | - | - |