
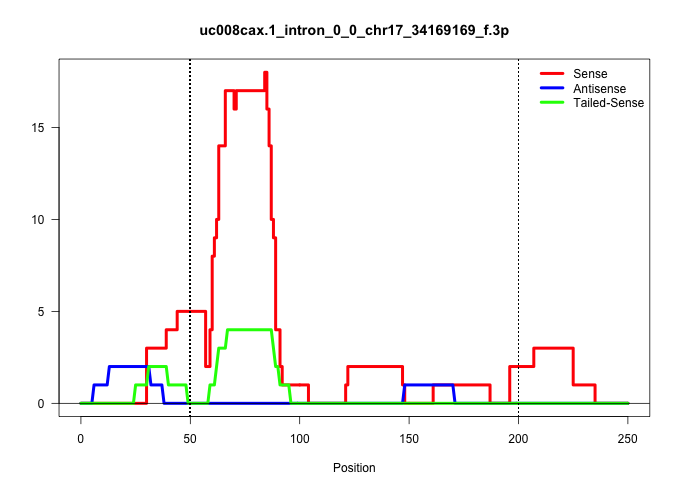
| Gene: Rxrb | ID: uc008cax.1_intron_0_0_chr17_34169169_f.3p | SPECIES: mm9 |
 |
 |
|
(1) OTHER.mut |
(1) OVARY |
(5) PIWI.ip |
(2) PIWI.mut |
(1) TDRD1.ip |
(20) TESTES |
| CCCTTTTTGTCAACCGCCTCCTCTCACGACTATTAGAAGTGAAATTGTTGTTTGGCCACTTGGTATTTCTGAACTCCAGGGTTGCGGGCTGTGGTGGGTCAGGGAGAATTAAAGTGTCCATTTGGATGATCAAAAGGCTCACTTTCTCTTCTTGTCTCTCCTGTCCGTGTGTGTCTGCCTTGTACGTACCCACTTCTCAGATTCCCGAAGCCCAGACAGCTCCTCCCCAAATCCCCTTTCTCAGGGGATC .............................................................(((((..........(((((((((((((...((.(((.((((((((...((((((.((.((((......)))).))..)))))).)))))))).))).)).))))))).......))))))...)))))............................................................ ..........................................................59..................................................................................................................................191......................................................... |
Size | Perfect hit | Total Norm | Perfect Norm | SRR248524(GSM733812) cell type: Thy1+ spermatogonial stem cellstra. (testes) | mjTestesWT1() Testes Data. (testes) | SRR028732(GSM400969) Mili-Tdrd1 KO associated. (mili testes) | SRR248523(GSM733811) cell type: Thy1+ spermatogonial stem cellstra. (testes) | SRR034119(GSM466729) Mili IP_Tdrd9+/- replicate2. (mili testes) | mjTestesWT2() Testes Data. (testes) | mjTestesWT3() Testes Data. (testes) | GSM509280(GSM509280) small RNA cloning by length. (testes) | SRR034118(GSM466728) Mili IP_Tdrd9+/- replicate1. (mili testes) | SRR034120(GSM466730) Mili IP_Tdrd9-/- replicate1. (mili testes) | SRR014230(GSM319954) 10 dpp Dnmt3L-KO MILI. (mili testes) | SRR034121(GSM466731) Mili IP_Tdrd9-/- replicate2. (mili testes) | SRR042486(GSM539878) mouse ovaries [09-002]. (ovary) | SRR014229(GSM319953) 10 dpp MILI. (mili testes) | SRR069809(GSM610965) small RNA sequencing; sample 1. (testes) | mjTestesKO8() Testes Data. (Zcchc11 testes) | SRR248527(GSM733815) cell type: spermatogonial stem cell enriched . (testes) | SRR014235(GSM319959) 2 dpp total. (testes) | SRR037901(GSM510437) testes_rep2. (testes) | SRR248526(GSM733814) cell type: Thy1- spermatogonial stem cellstra. (testes) | SRR028730(GSM400967) Tdrd1-associated. (tdrd1 testes) |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| .........................................................................................................................................................................TGTGTCTGCCTTGTACGTACCCACTTCTCAG.................................................. | 31 | 1 | 6.00 | 6.00 | - | 6.00 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - |
| ...............................................................TATTTCTGAACTCCAGGGTTGCGGGC................................................................................................................................................................. | 26 | 1 | 4.00 | 4.00 | 3.00 | - | - | 1.00 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - |
| ..............................TATTAGAAGTGAAATTGTTGTTTGGCC................................................................................................................................................................................................. | 27 | 1 | 3.00 | 3.00 | 1.00 | - | 1.00 | - | - | - | - | - | 1.00 | - | - | - | - | - | - | - | - | - | - | - | - |
| .............................................................................AGGGTTGCGGGCTGTG............................................................................................................................................................. | 16 | 1 | 3.00 | 3.00 | - | - | - | - | - | 3.00 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - |
| ....................................................................................................................................................................................................TCAGATTCCCGAAGCCCAGACAGCTCCTC......................... | 29 | 1 | 2.00 | 2.00 | - | - | - | - | - | - | - | - | - | 1.00 | - | 1.00 | - | - | - | - | - | - | - | - | - |
| ...........................................................TTGGTATTTCTGAACTCCAGGGTTGC..................................................................................................................................................................... | 26 | 1 | 2.00 | 2.00 | - | - | - | - | - | - | 1.00 | - | - | - | - | - | - | - | - | - | - | 1.00 | - | - | - |
| ............................................................TGGTATTTCTGAACTCCAGGGTTGCGG................................................................................................................................................................... | 27 | 1 | 2.00 | 2.00 | - | - | 1.00 | - | - | - | - | - | - | 1.00 | - | - | - | - | - | - | - | - | - | - | - |
| ............................................................TGGTATTTCTGAACTCCAGGGTTGCG.................................................................................................................................................................... | 26 | 1 | 2.00 | 2.00 | - | - | - | - | 2.00 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - |
| ....................................................................................CGGGCTGTGGTGGGTCAGGG.................................................................................................................................................. | 20 | 1 | 2.00 | 2.00 | - | - | - | 1.00 | - | - | - | - | - | - | - | - | - | - | - | 1.00 | - | - | - | - | - |
| ...........................................................TTGGTATTTCTGAACTCCAGGGTTGCGGa.................................................................................................................................................................. | 29 | a | 1.00 | 1.00 | - | - | 1.00 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - |
| ........................................................................ACTCCAGGGTTGCGGGCTGTGG............................................................................................................................................................ | 22 | 1 | 1.00 | 1.00 | - | - | - | - | - | - | 1.00 | - | - | - | - | - | - | - | - | - | - | - | - | - | - |
| .....................................................................TGAACTCCAGGGTTGCGGGCTGTGGT........................................................................................................................................................... | 26 | 1 | 1.00 | 1.00 | - | - | - | - | - | - | 1.00 | - | - | - | - | - | - | - | - | - | - | - | - | - | - |
| ..................................................................TTCTGAACTCCAGGGTTGCGGGCTGT.............................................................................................................................................................. | 26 | 1 | 1.00 | 1.00 | 1.00 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - |
| .............................................................GGTATTTCTGAACTCCAGGGTTGC..................................................................................................................................................................... | 24 | 1 | 1.00 | 1.00 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | 1.00 | - | - | - | - |
| ............................................TTGTTGTTTGGCCACTTGGTATTTCT.................................................................................................................................................................................... | 26 | 1 | 1.00 | 1.00 | - | - | - | - | - | - | - | - | - | - | - | - | - | 1.00 | - | - | - | - | - | - | - |
| .........................................................................................................................TTGGATGATCAAAAGGCTCACTTTCT....................................................................................................... | 26 | 1 | 1.00 | 1.00 | - | - | - | - | - | - | - | - | - | - | 1.00 | - | - | - | - | - | - | - | - | - | - |
| .......................................................................AACTCCAGGGTTGCGGGCTG............................................................................................................................................................... | 20 | 1 | 1.00 | 1.00 | - | - | - | - | - | - | - | 1.00 | - | - | - | - | - | - | - | - | - | - | - | - | - |
| ...............................................................................................................................................................................................................AAGCCCAGACAGCTCCTCCCCAAATCCC............... | 28 | 1 | 1.00 | 1.00 | - | - | - | - | 1.00 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - |
| ...................................................................TCTGAACTCCAGGGTTGCGGGCTGTGGTt.......................................................................................................................................................... | 29 | t | 1.00 | 0.00 | 1.00 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - |
| ..............................................................GTATTTCTGAACTCCAGGGTTGCGGGCaa............................................................................................................................................................... | 29 | aa | 1.00 | 0.00 | - | - | - | - | 1.00 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - |
| ...................................................................................GCGGGCTGTGGTGGGTCAGGGA................................................................................................................................................. | 22 | 1 | 1.00 | 1.00 | - | - | - | - | - | - | - | - | - | - | - | - | 1.00 | - | - | - | - | - | - | - | - |
| ..................................................................TTCTGAACTCCAGGGTTGCGGGCTG............................................................................................................................................................... | 25 | 1 | 1.00 | 1.00 | - | - | - | 1.00 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - |
| ...............................................................TATTTCTGAACTCCAGGGTTGCGGG.................................................................................................................................................................. | 25 | 1 | 1.00 | 1.00 | - | - | - | - | - | 1.00 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - |
| .................................................................................................................................................................TGTCCGTGTGTGTCTGCCTTGTACGT............................................................... | 26 | 1 | 1.00 | 1.00 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | 1.00 |
| .........................ACGACTATTAGcggt.................................................................................................................................................................................................................. | 15 | cggt | 1.00 | 0.00 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | 1.00 | - |
| ...........................................................TTGGTATTTCTGAACTCCAGGGTTGCGG................................................................................................................................................................... | 28 | 1 | 1.00 | 1.00 | - | - | 1.00 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - |
| .......................................TGAAATTGTTGTTTGGCCACTTGGTATT....................................................................................................................................................................................... | 28 | 1 | 1.00 | 1.00 | - | - | - | - | 1.00 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - |
| ...............................................................TATTTCTGAACTCCAGGGTTGCGGGa................................................................................................................................................................. | 26 | a | 1.00 | 1.00 | 1.00 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - |
| ..................................................................TTCTGAACTCCAGGGTTGCGGG.................................................................................................................................................................. | 22 | 1 | 1.00 | 1.00 | - | - | - | 1.00 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - |
| ...................................................................TCTGAACTCCAGGGTTGCGGGC................................................................................................................................................................. | 22 | 1 | 1.00 | 1.00 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | 1.00 | - | - | - | - | - | - |
| ...............................ATTAGAAGTGAAATctgt......................................................................................................................................................................................................... | 18 | ctgt | 1.00 | 0.00 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | 1.00 | - | - |
| ..........................................................................................................................TGGATGATCAAAAGGCTCACTTTCTC...................................................................................................... | 26 | 1 | 1.00 | 1.00 | - | - | - | 1.00 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - |
| ..............................................................GTATTTCTGAACTCCAGGGTTGCGG................................................................................................................................................................... | 25 | 1 | 1.00 | 1.00 | 1.00 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - |
| CCCTTTTTGTCAACCGCCTCCTCTCACGACTATTAGAAGTGAAATTGTTGTTTGGCCACTTGGTATTTCTGAACTCCAGGGTTGCGGGCTGTGGTGGGTCAGGGAGAATTAAAGTGTCCATTTGGATGATCAAAAGGCTCACTTTCTCTTCTTGTCTCTCCTGTCCGTGTGTGTCTGCCTTGTACGTACCCACTTCTCAGATTCCCGAAGCCCAGACAGCTCCTCCCCAAATCCCCTTTCTCAGGGGATC .............................................................(((((..........(((((((((((((...((.(((.((((((((...((((((.((.((((......)))).))..)))))).)))))))).))).)).))))))).......))))))...)))))............................................................ ..........................................................59..................................................................................................................................191......................................................... |
Size | Perfect hit | Total Norm | Perfect Norm | SRR248524(GSM733812) cell type: Thy1+ spermatogonial stem cellstra. (testes) | mjTestesWT1() Testes Data. (testes) | SRR028732(GSM400969) Mili-Tdrd1 KO associated. (mili testes) | SRR248523(GSM733811) cell type: Thy1+ spermatogonial stem cellstra. (testes) | SRR034119(GSM466729) Mili IP_Tdrd9+/- replicate2. (mili testes) | mjTestesWT2() Testes Data. (testes) | mjTestesWT3() Testes Data. (testes) | GSM509280(GSM509280) small RNA cloning by length. (testes) | SRR034118(GSM466728) Mili IP_Tdrd9+/- replicate1. (mili testes) | SRR034120(GSM466730) Mili IP_Tdrd9-/- replicate1. (mili testes) | SRR014230(GSM319954) 10 dpp Dnmt3L-KO MILI. (mili testes) | SRR034121(GSM466731) Mili IP_Tdrd9-/- replicate2. (mili testes) | SRR042486(GSM539878) mouse ovaries [09-002]. (ovary) | SRR014229(GSM319953) 10 dpp MILI. (mili testes) | SRR069809(GSM610965) small RNA sequencing; sample 1. (testes) | mjTestesKO8() Testes Data. (Zcchc11 testes) | SRR248527(GSM733815) cell type: spermatogonial stem cell enriched . (testes) | SRR014235(GSM319959) 2 dpp total. (testes) | SRR037901(GSM510437) testes_rep2. (testes) | SRR248526(GSM733814) cell type: Thy1- spermatogonial stem cellstra. (testes) | SRR028730(GSM400967) Tdrd1-associated. (tdrd1 testes) |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| .............CCGCCTCCTCTCACGACTATTAGAA.................................................................................................................................................................................................................... | 25 | 1 | 1.00 | 1.00 | - | - | 1.00 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - |
| ......TTGTCAACCGCCTCCTCTCACGACTA.......................................................................................................................................................................................................................... | 26 | 1 | 1.00 | 1.00 | - | - | - | - | - | - | - | - | 1.00 | - | - | - | - | - | - | - | - | - | - | - | - |
| ....................................................................................................................................................TTCTTGTCTCTCCTGTCCGTGTG............................................................................... | 23 | 1 | 1.00 | 1.00 | - | - | - | - | - | - | - | 1.00 | - | - | - | - | - | - | - | - | - | - | - | - | - |