
| Gene: BC024814 | ID: uc007ycl.1_intron_9_0_chr16_8488921_f.5p | SPECIES: mm9 |
 |
 |
|
(1) OTHER.mut |
(6) PIWI.ip |
(1) PIWI.mut |
(1) TDRD1.ip |
(24) TESTES |
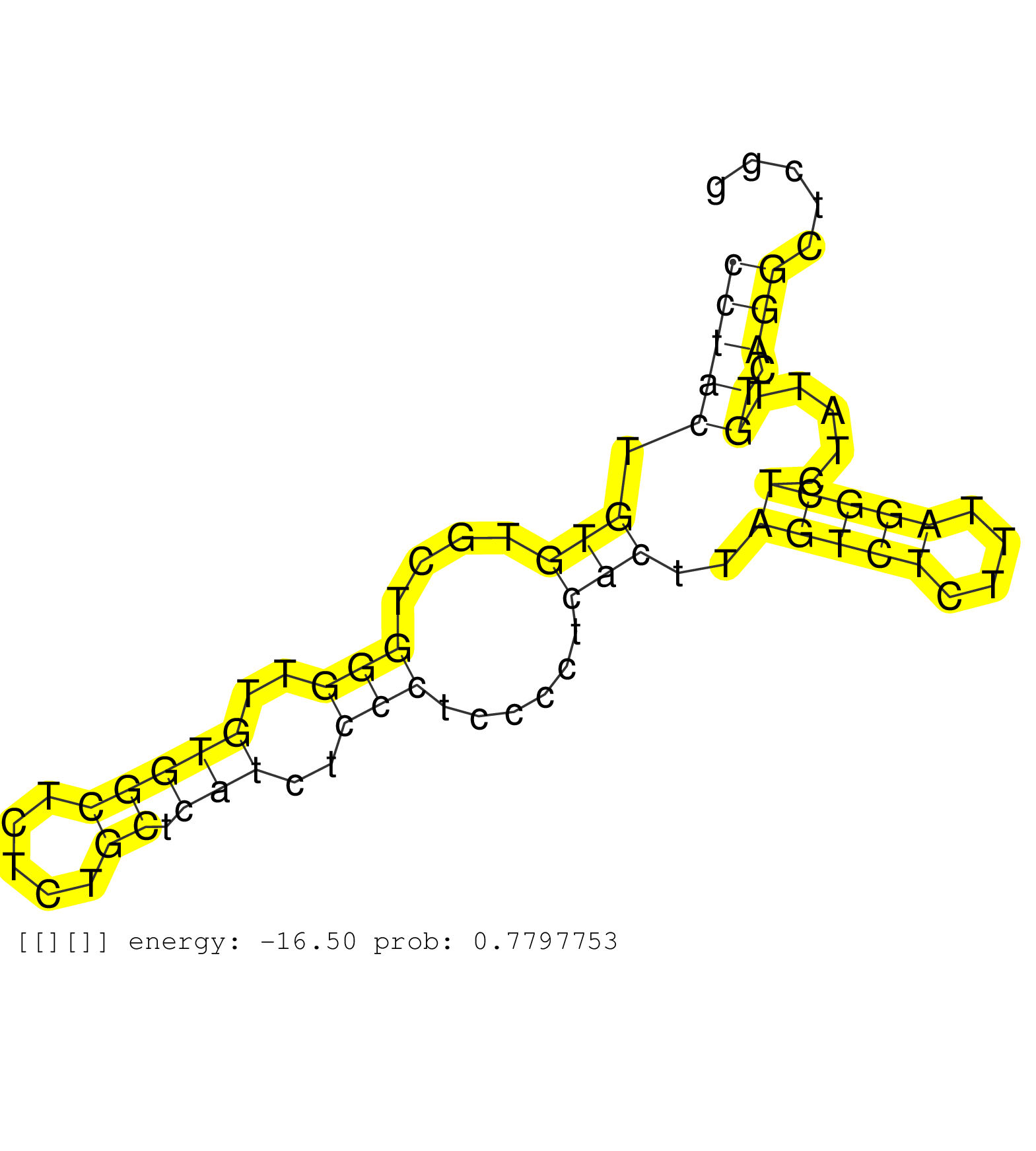
| TGGAGGCCTCCTTCCCGCAGCTGCTCGTCTATGAACGCATCCGGCAGCTGGTAGATGCAGCCCCGTGAGGGTGGTGGGAAGCAAGGCACTGGACATGTCACACACCTACTGTGTGCTGGGTTGTGGCTCTCTGCTCATCTCCCTCCCCTCACTTAGTCTCTTTAGGCTCTATTGTCAGGCTCGGCCAGCCTACTGTCCATCCATGGTCTGCCCGGGAGACAGAGACGCTGCAGGCCTAACACTGTGATCC ........................................................................................................(((((.(((....(((..(((((.....)).)))..)))......)))..(((((....))))).....)).)))....................................................................... ........................................................................................................105............................................................................184................................................................ |
Size | Perfect hit | Total Norm | Perfect Norm | SRR034119(GSM466729) Mili IP_Tdrd9+/- replicate2. (mili testes) | SRR034118(GSM466728) Mili IP_Tdrd9+/- replicate1. (mili testes) | SRR034121(GSM466731) Mili IP_Tdrd9-/- replicate2. (mili testes) | SRR248523(GSM733811) cell type: Thy1+ spermatogonial stem cellstra. (testes) | SRR034120(GSM466730) Mili IP_Tdrd9-/- replicate1. (mili testes) | SRR248524(GSM733812) cell type: Thy1+ spermatogonial stem cellstra. (testes) | SRR014236(GSM319960) 10 dpp total. (testes) | SRR014231(GSM319955) 16.5 dpc total. (testes) | SRR028732(GSM400969) Mili-Tdrd1 KO associated. (mili testes) | SRR069809(GSM610965) small RNA sequencing; sample 1. (testes) | mjTestesWT2() Testes Data. (testes) | SRR028731(GSM400968) Mili-wt-associated. (testes) | SRR069811(GSM610967) small RNA sequencing; sample 3. (testes) | SRR029038(GSM433290) 25dpp_hetero_tdrd6-KO. (tdrd6 testes) | GSM475280(GSM475280) Mili-IP. (mili testes) | SRR248527(GSM733815) cell type: spermatogonial stem cell enriched . (testes) | SRR014235(GSM319959) 2 dpp total. (testes) | SRR248526(GSM733814) cell type: Thy1- spermatogonial stem cellstra. (testes) | SRR014232(GSM319956) 16.5 dpc MILI. (mili testes) | SRR363957(GSM822759) P20-WTSmall RNA Miwi IPread_length: 36. (testes) | SRR248525(GSM733813) cell type: Thy1- spermatogonial stem cellstra. (testes) | SRR028730(GSM400967) Tdrd1-associated. (tdrd1 testes) | GSM509275(GSM509275) MitoPLD+/+ E16.5 small RNA. (testes) | GSM509279(GSM509279) MVH-/- E16.5 small RNA. (testes) |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| ...............................TGAACGCATCCGGCAGCTGGTAGAT.................................................................................................................................................................................................. | 25 | 1 | 18.00 | 18.00 | 4.00 | 7.00 | 3.00 | 1.00 | - | - | - | - | - | 1.00 | - | - | 1.00 | - | - | - | - | - | - | - | - | 1.00 | - | - |
| .....................TGCTCGTCTATGAACGCATCCGGCAGC.......................................................................................................................................................................................................... | 27 | 1 | 16.00 | 16.00 | 3.00 | 3.00 | 2.00 | - | 1.00 | - | - | - | 1.00 | 2.00 | 2.00 | - | - | - | - | - | - | - | 1.00 | - | - | - | - | 1.00 |
| .............................TATGAACGCATCCGGCAGCTGGTAGA................................................................................................................................................................................................... | 26 | 1 | 15.00 | 15.00 | 5.00 | 4.00 | - | 1.00 | 1.00 | 2.00 | - | - | 1.00 | - | - | - | 1.00 | - | - | - | - | - | - | - | - | - | - | - |
| ...........................TCTATGAACGCATCCGGCAGCTGGTA..................................................................................................................................................................................................... | 26 | 1 | 11.00 | 11.00 | 5.00 | 4.00 | - | - | 1.00 | - | 1.00 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - |
| .........................................................................................................................................................TAGTCTCTTTAGGCTCTATTGTCAGGC...................................................................... | 27 | 1 | 8.00 | 8.00 | 1.00 | 2.00 | 1.00 | 1.00 | 2.00 | 1.00 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - |
| ...............................TGAACGCATCCGGCAGCTGGTAGATG................................................................................................................................................................................................. | 26 | 1 | 8.00 | 8.00 | 3.00 | 2.00 | 1.00 | 2.00 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - |
| .....................TGCTCGTCTATGAACGCATCCGGCAG........................................................................................................................................................................................................... | 26 | 1 | 8.00 | 8.00 | 2.00 | 2.00 | 2.00 | - | 1.00 | - | - | - | - | 1.00 | - | - | - | - | - | - | - | - | - | - | - | - | - | - |
| ...............................TGAACGCATCCGGCAGCTGGTAGATt................................................................................................................................................................................................. | 26 | t | 6.00 | 18.00 | 1.00 | 4.00 | - | - | 1.00 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - |
| ................................................................................................................................................................................................................TGCCCGGGAGACAGAGACGCTGCAGGCCT............. | 29 | 1 | 6.00 | 6.00 | 4.00 | 1.00 | 1.00 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - |
| .............................................................................................................TGTGTGCTGGGTTGTGGCTCTCTGC.................................................................................................................... | 25 | 1 | 5.00 | 5.00 | - | - | - | 2.00 | - | 2.00 | - | - | - | - | 1.00 | - | - | - | - | - | - | - | - | - | - | - | - | - |
| .............................TATGAACGCATCCGGCAGCTGGTAGAT.................................................................................................................................................................................................. | 27 | 1 | 5.00 | 5.00 | 1.00 | 1.00 | 1.00 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | 1.00 | - | - | - | - | 1.00 | - |
| ................................................................................................................................................................................................................TGCCCGGGAGACAGAGACGCTGCAGGCC.............. | 28 | 1 | 4.00 | 4.00 | 3.00 | 1.00 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - |
| ................................................................................................................................................................................................................TGCCCGGGAGACAGAGACGCTGCAGGC............... | 27 | 1 | 3.00 | 3.00 | 1.00 | 2.00 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - |
| ........................TCGTCTATGAACGCATCCGGCAGCTGG....................................................................................................................................................................................................... | 27 | 1 | 3.00 | 3.00 | 3.00 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - |
| ..............................................................................................................................................................................................................TCTGCCCGGGAGACAGAGACGCTGCA.................. | 26 | 1 | 3.00 | 3.00 | 1.00 | 1.00 | 1.00 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - |
| ...........................TCTATGAACGCATCCGGCAGCTGGagc.................................................................................................................................................................................................... | 27 | agc | 3.00 | 0.00 | 2.00 | - | 1.00 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - |
| .............................................................................................................TGTGTGCTGGGTTGTGGCTCTCTGa.................................................................................................................... | 25 | a | 2.00 | 1.00 | - | - | - | - | - | - | 1.00 | - | - | - | - | - | - | - | - | - | 1.00 | - | - | - | - | - | - | - |
| ..................AGCTGCTCGTCTATGAACGCATCCGGC............................................................................................................................................................................................................. | 27 | 1 | 2.00 | 2.00 | - | 2.00 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - |
| .............................TATGAACGCATCCGGCAGCTGGTAG.................................................................................................................................................................................................... | 25 | 1 | 2.00 | 2.00 | 1.00 | - | - | 1.00 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - |
| .......................CTCGTCTATGAACGCATCCGGCAGCTGt....................................................................................................................................................................................................... | 28 | t | 2.00 | 1.00 | - | 1.00 | 1.00 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - |
| ...................................................................................................................................................................................................................CCGGGAGACAGAGACGCTGCAGGCCT............. | 26 | 1 | 2.00 | 2.00 | 1.00 | 1.00 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - |
| ................................................................................................................................................................TTTAGGCTCTATTGTCAGGCTCGGC................................................................. | 25 | 1 | 2.00 | 2.00 | - | - | - | 2.00 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - |
| .................................................................................................................................................................TTAGGCTCTATTGTCAGGCTCGGCCAGC............................................................. | 28 | 1 | 2.00 | 2.00 | - | - | - | - | 2.00 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - |
| ...........................TCTATGAACGCATCCGGCAGCTGGag..................................................................................................................................................................................................... | 26 | ag | 2.00 | 0.00 | 1.00 | - | - | - | - | - | - | - | - | - | - | 1.00 | - | - | - | - | - | - | - | - | - | - | - | - |
| ..........................................................................................................TACTGTGTGCTGGGTTGTGGCTCTC....................................................................................................................... | 25 | 1 | 2.00 | 2.00 | - | - | - | - | - | 1.00 | - | 1.00 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - |
| .........................................................................................................................................................TAGTCTCTTTAGGCTCTATTGTCAGGCT..................................................................... | 28 | 1 | 1.00 | 1.00 | - | - | - | - | 1.00 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - |
| ....................CTGCTCGTCTATGAACGCATCCGGCA............................................................................................................................................................................................................ | 26 | 1 | 1.00 | 1.00 | 1.00 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - |
| .............................................................................................................TGTGTGCTGGGTTGTGGCTCTCTG..................................................................................................................... | 24 | 1 | 1.00 | 1.00 | - | - | - | - | - | - | - | 1.00 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - |
| .....................TGCTCGTCTATGAACGCATCCGGCAGCTG........................................................................................................................................................................................................ | 29 | 1 | 1.00 | 1.00 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | 1.00 | - | - | - | - | - | - | - | - |
| ...............................TGAACGCATCCGGCAGCTGGTAGt................................................................................................................................................................................................... | 24 | t | 1.00 | 0.00 | - | - | - | 1.00 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - |
| ...........................TCTATGAACGCATCCGGCAGCTGGaga.................................................................................................................................................................................................... | 27 | aga | 1.00 | 0.00 | - | - | - | - | - | - | - | - | 1.00 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - |
| ................................................................................................................................................................................................................TGCCCGGGAGACAGAGACGCTGCAGGCa.............. | 28 | a | 1.00 | 3.00 | - | - | 1.00 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - |
| ...........TTCCCGCAGCTGCTCGTCTATGAACGC.................................................................................................................................................................................................................... | 27 | 1 | 1.00 | 1.00 | - | - | - | - | - | - | - | - | - | - | - | - | - | 1.00 | - | - | - | - | - | - | - | - | - | - |
| ...............................TGAACGCATCCGGCAGCTGGTAGA................................................................................................................................................................................................... | 24 | 1 | 1.00 | 1.00 | - | - | - | - | - | 1.00 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - |
| ................................................TGGTAGATGCAGCCCCGTGAGGGTGGTG.............................................................................................................................................................................. | 28 | 1 | 1.00 | 1.00 | - | - | - | - | - | - | 1.00 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - |
| .............................................................................................................TGTGTGCTGGGTTGTGGCTCTCTGCTt.................................................................................................................. | 27 | t | 1.00 | 1.00 | - | - | - | - | - | - | - | 1.00 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - |
| .............................TATGAACGCATCCGGC............................................................................................................................................................................................................. | 16 | 1 | 1.00 | 1.00 | - | - | - | 1.00 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - |
| ..............................................................CCGTGAGGGTGGTGGGAAGCAAGGCA.................................................................................................................................................................. | 26 | 1 | 1.00 | 1.00 | - | 1.00 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - |
| .....................TGCTCGTCTATGAACGCATCCGGCAGCTGG....................................................................................................................................................................................................... | 30 | 1 | 1.00 | 1.00 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | 1.00 | - | - | - | - |
| ........................................................................................................................................................TTAGTCTCTTTAGGCTCTATTGTCAGG....................................................................... | 27 | 1 | 1.00 | 1.00 | 1.00 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - |
| .......................CTCGTCTATGAACGCATCCGGCAGCTG........................................................................................................................................................................................................ | 27 | 1 | 1.00 | 1.00 | - | 1.00 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - |
| .............................................................................................................TGTGTGCTGGGTTGTGGCTCTCcgct................................................................................................................... | 26 | cgct | 1.00 | 0.00 | - | - | - | - | - | 1.00 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - |
| ................................................TGGTAGATGCAGCCCCGTGAGGGgggt............................................................................................................................................................................... | 27 | gggt | 1.00 | 0.00 | - | - | - | 1.00 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - |
| .............................................................................................................TGTGTGCTGGGTTGTGGCTCTCTGCTC.................................................................................................................. | 27 | 1 | 1.00 | 1.00 | - | - | - | - | - | - | - | 1.00 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - |
| ................................................................................................................................................................TTTAGGCTCTATTGTCAGGCTCGGCC................................................................ | 26 | 1 | 1.00 | 1.00 | 1.00 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - |
| .......................CTCGTCTATGAACGCATCCGGCAGCT......................................................................................................................................................................................................... | 26 | 1 | 1.00 | 1.00 | 1.00 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - |
| ...........................TCTATGAACGCATCCGGCAGCTGGTt..................................................................................................................................................................................................... | 26 | t | 1.00 | 0.00 | 1.00 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - |
| ....................................................................................................................................................................................................................CGGGAGACAGAGACGCTGCAGGCCTA............ | 26 | 1 | 1.00 | 1.00 | 1.00 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - |
| ........................................................................................................................................................TTAGTCTCTTTAGGCTCTATTGTCAGGC...................................................................... | 28 | 1 | 1.00 | 1.00 | - | - | - | - | - | - | - | - | 1.00 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - |
| .............................TATGAACGCATCCGGCAGCTGG....................................................................................................................................................................................................... | 22 | 1 | 1.00 | 1.00 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | 1.00 | - | - | - |
| ........TCCTTCCCGCAGCTGCTCGTCTATGAA....................................................................................................................................................................................................................... | 27 | 1 | 1.00 | 1.00 | 1.00 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - |
| ..................................................................................................................................................................................................................CCCGGGAGACAGAGACGCTGCAGGCCTt............ | 28 | t | 1.00 | 0.00 | - | - | - | - | 1.00 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - |
| .................................................................................................................................................................TTAGGCTCTATTGTCAGGCTCGGCCAGCC............................................................ | 29 | 1 | 1.00 | 1.00 | - | - | - | - | - | - | 1.00 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - |
| ................................................................................................................................................................TTTAGGCTCTATTGTCAGGCTCGGCCA............................................................... | 27 | 1 | 1.00 | 1.00 | 1.00 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - |
| ....................CTGCTCGTCTATGAACGCATCCGGCAG........................................................................................................................................................................................................... | 27 | 1 | 1.00 | 1.00 | - | - | - | - | - | - | - | - | - | - | - | 1.00 | - | - | - | - | - | - | - | - | - | - | - | - |
| ........................................................................................................CCTACTGTGTGCTGGGTTGTGGCTCT........................................................................................................................ | 26 | 1 | 1.00 | 1.00 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | 1.00 | - | - | - | - | - | - | - | - | - |
| .......................CTCGTCTATGAACGCATCCGGCAGC.......................................................................................................................................................................................................... | 25 | 1 | 1.00 | 1.00 | - | - | - | - | - | - | 1.00 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - |
| .............................................................................................................TGTGTGCTGGGTTGTGGCTCTCTGCT................................................................................................................... | 26 | 1 | 1.00 | 1.00 | - | - | - | - | - | - | - | 1.00 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - |
| ........................TCGTCTATGAACGCATCCGGCAGCTGGagt.................................................................................................................................................................................................... | 30 | agt | 1.00 | 3.00 | 1.00 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - |
| ....................................................................................................................................................................................................................CGGGAGACAGAGACGCTGCAGGCCTAA........... | 27 | 1 | 1.00 | 1.00 | - | 1.00 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - |
| .............................................................................................................TGTGTGCTGGGTTGTGG............................................................................................................................ | 17 | 2 | 0.50 | 0.50 | - | - | - | 0.50 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - |
| TGGAGGCCTCCTTCCCGCAGCTGCTCGTCTATGAACGCATCCGGCAGCTGGTAGATGCAGCCCCGTGAGGGTGGTGGGAAGCAAGGCACTGGACATGTCACACACCTACTGTGTGCTGGGTTGTGGCTCTCTGCTCATCTCCCTCCCCTCACTTAGTCTCTTTAGGCTCTATTGTCAGGCTCGGCCAGCCTACTGTCCATCCATGGTCTGCCCGGGAGACAGAGACGCTGCAGGCCTAACACTGTGATCC ........................................................................................................(((((.(((....(((..(((((.....)).)))..)))......)))..(((((....))))).....)).)))....................................................................... ........................................................................................................105............................................................................184................................................................ |
Size | Perfect hit | Total Norm | Perfect Norm | SRR034119(GSM466729) Mili IP_Tdrd9+/- replicate2. (mili testes) | SRR034118(GSM466728) Mili IP_Tdrd9+/- replicate1. (mili testes) | SRR034121(GSM466731) Mili IP_Tdrd9-/- replicate2. (mili testes) | SRR248523(GSM733811) cell type: Thy1+ spermatogonial stem cellstra. (testes) | SRR034120(GSM466730) Mili IP_Tdrd9-/- replicate1. (mili testes) | SRR248524(GSM733812) cell type: Thy1+ spermatogonial stem cellstra. (testes) | SRR014236(GSM319960) 10 dpp total. (testes) | SRR014231(GSM319955) 16.5 dpc total. (testes) | SRR028732(GSM400969) Mili-Tdrd1 KO associated. (mili testes) | SRR069809(GSM610965) small RNA sequencing; sample 1. (testes) | mjTestesWT2() Testes Data. (testes) | SRR028731(GSM400968) Mili-wt-associated. (testes) | SRR069811(GSM610967) small RNA sequencing; sample 3. (testes) | SRR029038(GSM433290) 25dpp_hetero_tdrd6-KO. (tdrd6 testes) | GSM475280(GSM475280) Mili-IP. (mili testes) | SRR248527(GSM733815) cell type: spermatogonial stem cell enriched . (testes) | SRR014235(GSM319959) 2 dpp total. (testes) | SRR248526(GSM733814) cell type: Thy1- spermatogonial stem cellstra. (testes) | SRR014232(GSM319956) 16.5 dpc MILI. (mili testes) | SRR363957(GSM822759) P20-WTSmall RNA Miwi IPread_length: 36. (testes) | SRR248525(GSM733813) cell type: Thy1- spermatogonial stem cellstra. (testes) | SRR028730(GSM400967) Tdrd1-associated. (tdrd1 testes) | GSM509275(GSM509275) MitoPLD+/+ E16.5 small RNA. (testes) | GSM509279(GSM509279) MVH-/- E16.5 small RNA. (testes) |
|---|